EEG functional source localisation
|
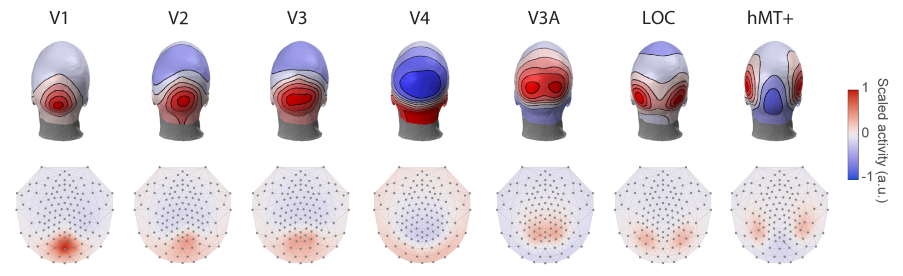
Together with Dr Justin Ales, I developed a very exciting method for localising brain sources from EEG recordings (Poncet & Ales, 2023). It is accurate, inexpensive, rapid, and easy-to-use (no need to install special software or collect and analyse MRI scans). Briefly, the method uses topographies of EEG activity derived for multiple functionally defined visual brain area to recover the intracranial sources responsible for a given EEG scalp activity. Thus, an important aspect of this method is that instead of recovering anatomical hard-to-interpret brain sources, it recovers functional sources. This makes the results easily comparable across EEG experiments (even if they use different setups) but also with MEG, fMRI or intracranial recordings.
|
The implementation of this method is available at https://github.com/aleslab/eegSourceTemplateMatching. It is compatible across all EEG systems (Biosemi, EGI, Easycap, Neuroscan, etc., with any number of electrodes; it only needs information about the electrode locations) and EEG software (EEGlab, Fieldtrip or user-based).
Goodness-of-fit measure for EEG
In our study (Poncet & Ales, 2020), we compared predicted and recorded EEG responses in different conditions. For example, we compared the recorded EEG response when an object was presented successively on the left and right of the screen and perceived as moving with the predicted EEG response created from the sum of the EEG response for the object presented only on the left or only on the right (with no perceived motion).
To test how well the prediction fits the recorded EEG, one can compute the amount of difference between the two signals using the root-mean-square error (RMSE). A major challenge with this comparison is to determine whether the resulting difference is a real signal that is not accounted by the prediction, or if it is just noise. For this, we developed a novel mathematical formulation that takes into consideration the variance of the noise in the recorded and predicted signal. With this quantification we can determine an objective level of noise that can be compared to the difference between the recorded and predicted signal. The closer the two values are, the better the prediction is to the recorded EEG.
For more details, see the paragraph “Prediction Error” in the Method section and equations 9 to 13.
To test how well the prediction fits the recorded EEG, one can compute the amount of difference between the two signals using the root-mean-square error (RMSE). A major challenge with this comparison is to determine whether the resulting difference is a real signal that is not accounted by the prediction, or if it is just noise. For this, we developed a novel mathematical formulation that takes into consideration the variance of the noise in the recorded and predicted signal. With this quantification we can determine an objective level of noise that can be compared to the difference between the recorded and predicted signal. The closer the two values are, the better the prediction is to the recorded EEG.
For more details, see the paragraph “Prediction Error” in the Method section and equations 9 to 13.